PK/PD, Exposure-Response - Time to Event
Rmarkdown template to generate this page can be found on Rmarkdown-Template.
Explore Exposure-Response Relationship
Time-to-event plots can be summarized by Kaplan-Meier plots and stratified by dose to give an overview of the dose-response
km_data <- lung %>%
mutate(fake_exposure = ph.ecog + age/50) %>%
filter(!is.na(fake_exposure)) %>%
mutate(expos_quantile= cut(fake_exposure,
breaks=quantile(fake_exposure,c(0,.25,.5,.75,1)),
include.lowest=TRUE,
labels=paste0("Q",c(4,3,2,1))))
km_fit <- survfit(Surv(time) ~ expos_quantile, data = km_data)
km_fit2 <- summary(km_fit, times = c(1,30,60,90*(1:10)))
data = data.frame(time =km_fit2$time,
surv= km_fit2$surv,
exposure_quantile = km_fit2$strata) %>%
mutate(exposure_quantile = stringr::str_replace(exposure_quantile,"expos_quantile=",""))
gg <- ggplot(data, aes(x=time, y=surv, color = exposure_quantile))
gg <- gg + geom_line(size = 1)
gg <- gg + scale_y_continuous(labels=scales::percent_format())
gg <- gg + ylab("Survival") + xlab("Time")
gg <- gg + guides(color=guide_legend(""))
gg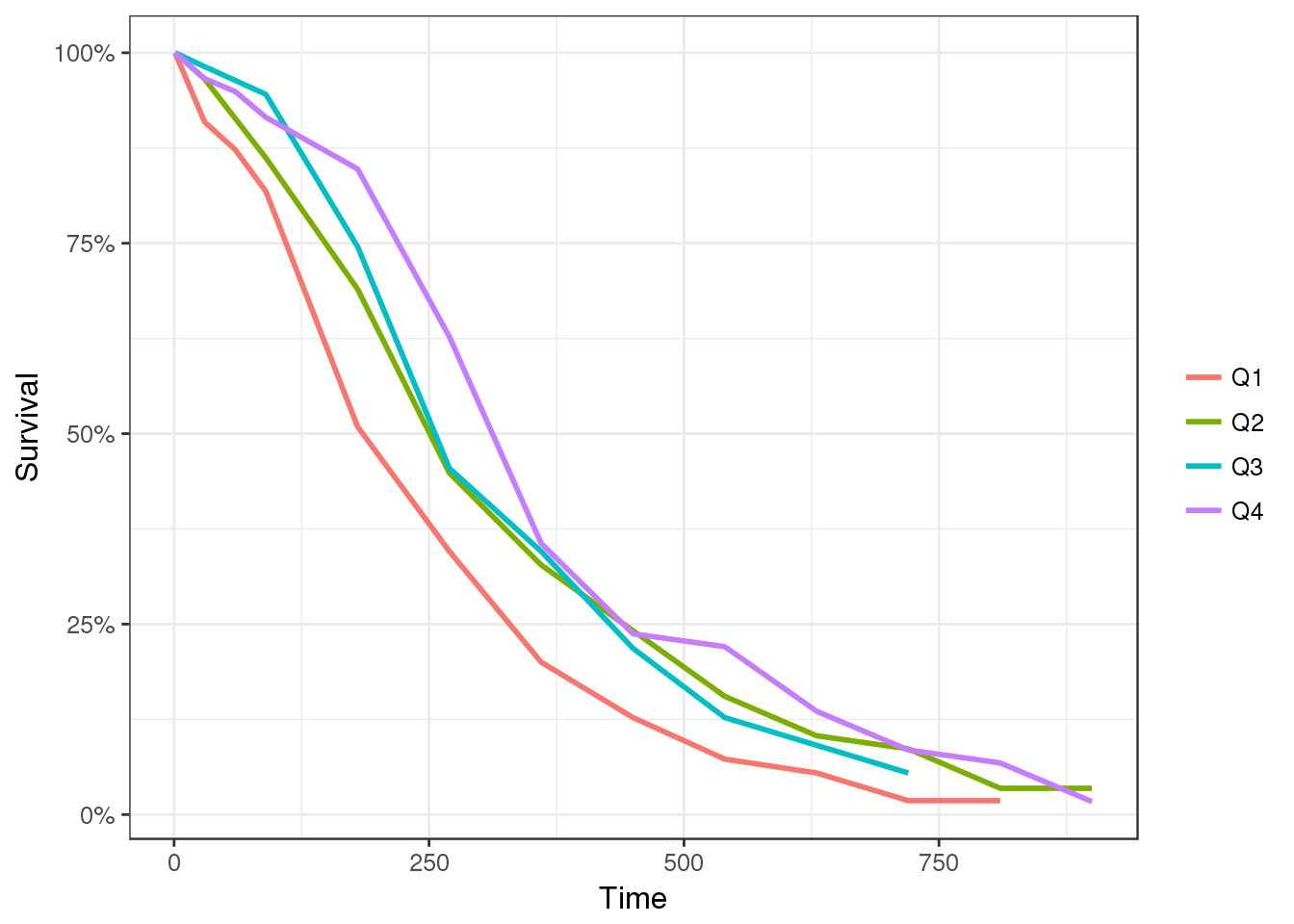
R Session Info
sessionInfo()## R version 3.4.3 (2017-11-30)
## Platform: x86_64-pc-linux-gnu (64-bit)
## Running under: Red Hat Enterprise Linux Server 7.4 (Maipo)
##
## Matrix products: default
## BLAS/LAPACK: /CHBS/apps/intel/17.4.196/compilers_and_libraries_2017.4.196/linux/mkl/lib/intel64_lin/libmkl_gf_lp64.so
##
## locale:
## [1] LC_CTYPE=en_US.UTF-8 LC_NUMERIC=C
## [3] LC_TIME=en_US.UTF-8 LC_COLLATE=en_US.UTF-8
## [5] LC_MONETARY=en_US.UTF-8 LC_MESSAGES=en_US.UTF-8
## [7] LC_PAPER=en_US.UTF-8 LC_NAME=C
## [9] LC_ADDRESS=C LC_TELEPHONE=C
## [11] LC_MEASUREMENT=en_US.UTF-8 LC_IDENTIFICATION=C
##
## attached base packages:
## [1] grid stats graphics grDevices utils datasets methods
## [8] base
##
## other attached packages:
## [1] reshape_0.8.7 lubridate_1.7.1 survival_2.41-3 DT_0.2
## [5] RxODE_0.6-1 bindrcpp_0.2 haven_1.1.0 readr_1.1.1
## [9] readxl_1.0.0 xtable_1.8-2 tidyr_0.7.2 caTools_1.17.1
## [13] zoo_1.8-0 dplyr_0.7.4 ggplot2_2.2.1 gridExtra_2.3
##
## loaded via a namespace (and not attached):
## [1] purrr_0.2.4 reshape2_1.4.3 splines_3.4.3
## [4] lattice_0.20-35 colorspace_1.3-2 htmltools_0.3.6
## [7] yaml_2.1.16 rlang_0.1.6 pillar_1.0.1
## [10] glue_1.2.0 RColorBrewer_1.1-2 binom_1.1-1
## [13] bindr_0.1 plyr_1.8.4 stringr_1.2.0
## [16] munsell_0.4.3 gtable_0.2.0 cellranger_1.1.0
## [19] htmlwidgets_0.9 codetools_0.2-15 evaluate_0.10.1
## [22] memoise_1.1.0 labeling_0.3 knitr_1.18
## [25] forcats_0.2.0 rex_1.1.2 markdown_0.8
## [28] Rcpp_0.12.14 scales_0.5.0 backports_1.1.2
## [31] jsonlite_1.5 hms_0.4.0 digest_0.6.13
## [34] stringi_1.1.3 rprojroot_1.3-1 tools_3.4.3
## [37] bitops_1.0-6 magrittr_1.5 lazyeval_0.2.1
## [40] tibble_1.4.1 pkgconfig_2.0.1 Matrix_1.2-12
## [43] rsconnect_0.8.5 assertthat_0.2.0 rmarkdown_1.8
## [46] R6_2.2.2 compiler_3.4.3